coliwordpress安装
wordpress安装 时间:2021-01-12 阅读:()
www.
454.
com454SequencingTheRoadtotheFutureNovember13,2012IMPORTANTNOTICEIntendedUseUnlessexplicitlystatedotherwise,allRocheAppliedScienceand454LifeSciencesproductsandservicesreferencedinthispresentation/documentareintendedforthefollowinguse:Forlifescienceresearchonly.
Notforuseindiagnosticprocedures.
454SequencingReadLengthEvolutionRaisingthebarinnext-gensequencingGS20GSFLXStandard100bp250bp200520072008GSFLXTitanium400bp2011Upto1,000bpGSFLX+New!
ReadLength(bp)1,000500250750www.
454.
comGSFLX+SystemOverviewLatestgenerationoftheGSFLXSystemAvailableasnewinstrumentorupgradetoexistinginstrumentNewGSFLXTitaniumSequencingKitXL+forextra-longreadsequencing.
UsesexistingRapidLibraryPrepandemPCRkitsBackwardscompatiblewithexistingGSFLXTitaniumSequencingKitXLR70NewGSFLX+ComputingStation:4GSJuniorSystemWidelypublishedNGSbenchtopplatformWholeMicrobialGenomeSequencingMetagenomicsTranscriptomeSequencingGenotypingUltra-DeepSequencingReadthecompletelistofpeer-reviewedpublicationsatwww.
454.
com/publicationsOnereadOnewellOnebeadOnefragmentOneFragment,OneBead,OneReadShearDNAandaddlinkers'EmulsionPCR'ClonalamplificationDepositionofbeadsintowellsofPTPdeviceSequencing-by-synthesisDetectionofPPireleasewww.
454.
comA)Single-strandedtemplatemixedwithCaptureBeadsB)EmulsifymillionsofbeadsinPCRreagentstoformwater-in-oilmicroreactorsMicroreactorcontainscompleteamplificationmixC)AmplifyD)BreakMicroreactorsE)EnrichforDNApositiveBeforePCRAfterPCRGSFLX+ChemistryEmulsionPCRBasesareflowedsequentiallyandinaspecifiedcycleorder(TACG)AnucleotidecomplementarytothetemplatestrandgenerateslightThelightsignalisrecordedbytheCCDcameraduringeachflowSignalintensityisproportionaltothenumberofnucleotidesincorporatedMassivelyparallelsequencingofclonallyamplifiedbeadsinpicoliter-sizewells454SequencingSystemsKeyConceptwww.
454.
comSequencingBySynthesisAATCGGCATGCTAAAACTGARepeateddNTPflowsequence:AAGCTAAnnealPrimerProcesscontinuesuntiluser-definednumberofnucleotideflowcyclesarecompleted.
GSFLX+ChemistrySequencing-by-synthesisGCTATTTGCCGCGACGTTTTTADataProcessingGSRunProcessorTAGCTImageProcessingRegiondefinition&backgroundsubtractionWellidentificationonthePTPdeviceSignalProcessingNormalization&correctionstepsQualityfiltering&qualityreadtrimmingFlowgramgeneration&basecallingTACGFlowOrderKeysequenceTCAGorGACT(libraryreads)KeysequenceCATGorATGC(controlreads)UsedforwellidentificationandsignalcalibrationTTCTGCGAA454SequencingSystemsFlowgramGeneration&Basecallingwww.
454.
comDataAnalysisOverview(SFFTools)–SFFfilemanipulationtoolsGSdenovoAssembler–GenomeorTranscriptomedenovoassembly;Largegenomesuptohumansize;Hybridassemblies–Pairwisealignmentsofreadsresultinginconsensusandcorrespondingqualityscoresofcontigs,ACEfileofthealignment,metricsfilesandcontigscaffolds(whenusingPaired-enddata)GSReferenceMapper–TargetedmappingusingSequenceCapture;GenomeorTranscriptomereferencemapping;Variationdetectionincludinglargerearrangements–Individualreadsmappedtoareferencesequenceresultinginconsensusandcorrespondingqualityscoreofcontigs,ACEfileofthealignment,metricsfilesandlistofvariants(differencebetweenthesampleconsensusandreference)andstructuralrearrangements.
GSAmpliconVariantAnalyzer–Ultra-deepsequencingforvariantanalyses;Haplotyping–IdentificationandquantitationofsequencevariantsinanWhyReadLength,Speed&QualityMatterSequencingakillerE.
colibugwww.
454.
comDeadlyE.
colioutbreakinEuropebeganinMay2011ScientistsfromvariousinstitutesandcompaniesaroundtheglobequicklysequencedstrainisolatesThosewithfirstaccesstosampleshadtheleadwithspeed,butwhatofdataqualityandreadlengthWiththeGSJuniorassembly,theexpertswereabletoanswerbiologically-relevantquestionswhereotherassembliesfailed.
www.
454.
comE.
coliO104OutbreakSequencingtimelineMay24June1June2June6June10Germanauthoritiesreportthreedeathsand80casesofhaemolyticuraemicsyndromefromE.
coliinfectionEurope'stophealthofficialannounces"aseriouscrisis"UKHealthProtectionAgencyreleasesdenovoassemblyofH112180280isolateusingGSJuniorSystemUniversityHospitalMuenster&IonTorrentannounce"assembly"ofLB226692isolateBeijingGenomicsInstitutereleasesassemblyofTY2428isolate10PGMruns7PGM+1HiSeqruns3GSJuniorrunswww.
454.
comE.
coliO104OutbreakSequencingtimelineMay24June1June2June9June10"Moresequencedata,particularlypaired-endsequencingandlongerreadsthanwehaveinthecurrentassemblyarereallyneededtoresolvetheseprophageregions.
.
.
"NicoPetty,UniversityofQueenslandCommunityembracesHPAassemblyasreference–answersbiologicallymeaningfulquestionsBioinformaticsCommunityHitsaWallSource:http://bacpathgenomics.
wordpress.
com/2011/06/09/german-e-coli-phage-analysis-by-nico-petty/Highqualitydraftassembly(13scaffolds)Lotsofquestions……answers!
Globalbioinformaticscommunityanalyzingshort-readdatasetsLotsofdatabutlimitedcontextHighlyfragmentedassembliesrangingfrom>450to>3,000contigsRecognizedoutbreakstraingenomewasdifferentbutcouldn'tarrangethepieces"Afteronly5minutesoflookingovertheHPAassemblyweveryclearlysawwhatwehadexpectedtoseeallalongbutwasmissinginthepreviousassemblies–a62KbphageinsertioneventinthewrbAgene.
"DavidHirschberg,ColumbiaUniversityBGIReleasesIonTorrent–IlluminaAssemblyLotsofdatabutfewanswersBGIAnalysisWorkflowwww.
454.
comComplicatedpost-assemblyanalysis(evenfortheexperts)Complex,manualassemblyprocessQuestionsremainWhyisthisstrainsotoxicSomepiecesdon'tmatchtheexpectedbiology!
(Isthatreal)Source:http://bacpathgenomics.
wordpress.
com/2011/06/09/german-e-coli-phage-analysis-by-nico-petty/HPAReleasesGSJuniorAssemblyCommunitystartsansweringquestionswww.
454.
comGSJuniorSequenceDataGeneratedDeNovoAssembly(GSAssembler)Push-buttondenovoassemblyPerformedbyhealthcarescientistatHPA–notabioinformatician!
Nowthestorystartstoform…LargeinsertioneventsAcquiredmulti-drugresistanceAdhesin(knownvirulencefactor)Integratedintothechromosome!
Source:http://bacpathgenomics.
wordpress.
com/2011/06/11/tn21-resistance-transposon-in-the-chromosome454WorkflowThebiologybecomesclear~1minuteset-up~20mincomputationAnswersinhoursStx-phagescarriesgenesforShigatoxinPhage(62kblong)istypicallyintegratedintothewrbAgeneBothshortreadassembliesfailedtoaccuratelyreconstructthisportionofthegenomeRepresentationofStx2phageinsertiononHPAGSJuniorscaffold1WrbA5'endWrbA3'endStx2convertingphageTheBiology:WhyLongQualityReadsMatterBiologicallyaccurateinsertionreconstructionGSJuniorassemblycorrectlycontainstheinterruptedwrbAgenewithstx2phageinsertedonasinglescaffoldasisexpectedfromthebiologyBlogTitle:"Readlengthmatters:identifyingthephiStx2attsite""AmultisequencealignmentofthewrbAgeneclearlyrevealsthattheLife-TechnologiesAFOB00000000assemblingisnotreliableattheattachmentsite.
ItcontainsthreecontigsmatchingthewrbAgene,twoofthem(AFOB01000188.
1,AFOB01000143.
1)representingtheinterruptedgene,whilethethirdcontig(AFOB01000030.
1)showsanuninterruptedgene(thoughwithadistortedreadingframe).
"AssemblyComparisonOverviewFewerruns,morecompleteassembly,betterbiologyHPAAssemblyvs.
BGIAssembly3GSJuniorruns7IonTorrent+unspecifiedHiSeqrunsSingle-stepassemblyComplexmanualexpertassembly13scaffolds(35Xfewer)453scaffolds969KbN50scaffold(8.
5Xlonger)115KbN50scaffold~50%ofassembledbasesinscaffolds>1Mb0%Note:WithoutIlluminadata,IonTorrent&www.
454.
comCostComparisonOverviewFewerruns,easieranalysis,lowercostHPAGSJuniorassembly–3runs=~$3,000–1instrument=$100,000–Totalsequencingruntime=~1day–Assemblysoftware=FREE–Bioinformaticsexpertise=molecularbiologistfriendlyBGI(IonTorrent&Illumina)assembly–7IonTorrentruns=~$3,500–UnknownHiSeqpairedendruns=$20,000–1IonTorrentPGM&1IlluminaHiSeqinstrument=~$750,000–Totalsequencingruntime=1week+–AssemblySoftware/bioinformaticsexpertise=$$Expensive!
www.
454.
comGSJuniorAssemblyOverviewDeliveringwhatthecommunityneededUponrelease,communityimmediatelyembracedHPAassemblyasthereferenceGSJuniorassemblyprovidedclearanswerstokeybiologicalquestionsregardingthepathogenicityofthisuniquestrain–Correctlyidentifiedandlocateddrugresistanceandtoxingenesintegrationintochromosome–whileotherassembliescouldn't–Denovoassembledmajorityofplasmidcontent–Correctedmis-assembledphageregionsintheshort-readassembliesAfterworkingwithshortreadassembliesfordays,expertsturnedtotheGSJuniordataandwithin~5minutesfoundtheanswersPush-buttondenovoassemblyBiologicalanswers3GSJuniorRuns454SequencingDevelopmentsImprovingeaseofuse,sequencingperformance,andapplicationsUpfrontWorkflowAutomationSequencingReadLengthImprovementsApplications&AssaysEaseofuseimprovementstoGSJuniorworkflowStep-by-stepautomationsolutionsforlibraryprepandemPCR(breaking&enrichment)ExtendGSJuniorreadlengthbeyond400bp(shotgun&licon)OptimizeGSFLX+longreadsforampliconseqImproverobustnessandconsistencyofresultsLaunchdedicatedassaysforHLA,leukemiaandHIV/HBV/HCVPlannedCE-IVDapprovedkitsforselectassaysGSGTypeAssaysfor454SequencingSystemsAvailableNow!
GSGTypeLeukemiaPrimerSets(TET2/CBL/KRAS&RUNX1)GSGTypeHLAMR&HRPrimerSetsSequence-basedassaysforhigh-andmedium-resolutionHLAtypingForuseoneithertheGSJuniororGSFLXSystemSequence-basedassaysforleukemiaresearchForuseoneithertheGSJuniororGSFLXSystemExpanded454AssayMenuBeyond2012–focusonvirologyassaysVirology–GSVTectHIVResistance(PlannedCE-IVDinEurope)–HCVResistance/Genotyping(PlannedCE-IVDinEurope)–HBVResistance/Genotyping(PlannedCE-IVDinEurope)HLA–GSGTypeHLAPrimerSets(LSR)-AvailableNow!
–GSGTypeHLARegistry(PlannedCE-IVDinEurope)CollaborationwithDNAElectronicsISFETSequencingTechnologyCollaborationtodevelopsemiconductor-basedsequencingsystemLicenseDNAe'sISFET(ion-sensitive,fieldeffecttransistor)sequencingtechnologyEnablesdetectionofpHchangesfromnucleotideincorporationonsemiconductorchipBuildsoncurrent454SequencingportfoliobymovingfromopticaltoelectrochemicaldetectionTargetapplicationsinampliconassays,genepanels,exomesequencing,RNA-seq,andwholeDNAElectronicsSpunoutofImperialCollegeLondonDescriptionHigh-densityarrayofmicro-wellsonanintegratedcircuit.
EachwellholdsauniqueDNAtemplatebead.
Beneatheachwellisanion-sensitivetransistor.
Thechipissequentiallyfloodedwithonenucleotideafteranother.
IfthenucleotidethatfloodsthechipisnotamatchtotheDNAtemplate,nochangewillbedetectedandnobasewillbecalled.
IfanucleotideisincorporatedintotheCollaborationwithDNAe:TechnologyISFETsenablelabelfreepHSensingonaSemiconductorChipTechnologyConceptDescriptionMicrowellArrayPackagedIntegratedCircuitH+ionsdetectedCollaborationwithIBMNanoporeSequencingTechnologyIBMT.
J.
WatsonResearchCenterDescriptionCollaborationwithIBMResearchtodevelopnanopore-basedsequencingsystemTechnologybasedonIBMDNATransistorconceptLeverageIBMasaworldleaderinmicroelectronics,informationtechnologyandcomputationalbiologyAccesstoworldclassprototypefacilitiesforsemiconductormanufacturingandnanofabricationIBMDNAtransistorconceptcomprisesoflayersofinterdigitatedelectrodeswithdielectricmaterial,builtintoamembranethatcontainsthenanopores.
AnelectricfieldisusedtomoveDNAacrossananoporeonebaseatatime(electrophoreticratchet).
TheelectricfieldwithinthenanoporeinteractswithelectricchargesonDNAbackbonewhereindividualnucleotidesaretrappedatlocalfieldminima.
SwitchingthepolarityofthevoltageacrosstheelectrodespermitsthedriftoftheDNAmoleculeinonedirectionuntilthenucleotidesgettrappedatnewlocalfieldminimaAnucleotidesensingmethodologybasedonrecognitiontunnelingfromASUwillbecoupledtotheDNAtransistortodeterminetheCollaborationwithIBM:TechnologyConceptCombiningadielectricnanoporetransistorwithanucleotidesensingtechnologytoidentifyDNAbasesDNATransistorTechnologyConceptDescriptionNanoporeLocalElectricFieldMinimaDNAWatchthetechnologyvideoat:http://www.
youtube.
com/watchv=wvclP3GySUY454SequencingDevelopmentsSummaryOurobjectiveisto:improve–GSJunior&GSFLX+Systemsstrengthen-Assaysgrow–DNAecollaborationinnovate–IBMcollaborationwww.
454.
comForlifescienceresearchonly.
Notforuseindiagnosticprocedures.
454,454SEQUENCING,454LIFESCIENCES,EMPCR,GS20,GSFLX,GSJUNIOR,GSFLXTITANIUM,PTP,FASTSTARTandPICOTITERPLATEaretrademarksofRoche.
RocheTechnicalSupportus.
gssupport@roche.
com1.
800.
262.
49112-23-12Version1AnotherRecentExampleSequencingStreptococcusinToxicScarletFeverOutbreakJune15:UniversityofHongKongusedGSFLXSystemtosequenceStreptococcusstrainthatcausedscarletfeveroutbreakandonereporteddeathDraftgenomefinishedwithin3days–GSFLXrun(514,422reads)assembledusingGSDeNovoAssembler–47contigs–81KbN50contigsizeSource:http://www.
hku.
hk/press/news_detail_6505.
htmlResearchersidentifiedauniquemutation(48Kbinsertion)withingenomethatisabsentinotherpublishedgenomesofrelatedstrains
454.
com454SequencingTheRoadtotheFutureNovember13,2012IMPORTANTNOTICEIntendedUseUnlessexplicitlystatedotherwise,allRocheAppliedScienceand454LifeSciencesproductsandservicesreferencedinthispresentation/documentareintendedforthefollowinguse:Forlifescienceresearchonly.
Notforuseindiagnosticprocedures.
454SequencingReadLengthEvolutionRaisingthebarinnext-gensequencingGS20GSFLXStandard100bp250bp200520072008GSFLXTitanium400bp2011Upto1,000bpGSFLX+New!
ReadLength(bp)1,000500250750www.
454.
comGSFLX+SystemOverviewLatestgenerationoftheGSFLXSystemAvailableasnewinstrumentorupgradetoexistinginstrumentNewGSFLXTitaniumSequencingKitXL+forextra-longreadsequencing.
UsesexistingRapidLibraryPrepandemPCRkitsBackwardscompatiblewithexistingGSFLXTitaniumSequencingKitXLR70NewGSFLX+ComputingStation:4GSJuniorSystemWidelypublishedNGSbenchtopplatformWholeMicrobialGenomeSequencingMetagenomicsTranscriptomeSequencingGenotypingUltra-DeepSequencingReadthecompletelistofpeer-reviewedpublicationsatwww.
454.
com/publicationsOnereadOnewellOnebeadOnefragmentOneFragment,OneBead,OneReadShearDNAandaddlinkers'EmulsionPCR'ClonalamplificationDepositionofbeadsintowellsofPTPdeviceSequencing-by-synthesisDetectionofPPireleasewww.
454.
comA)Single-strandedtemplatemixedwithCaptureBeadsB)EmulsifymillionsofbeadsinPCRreagentstoformwater-in-oilmicroreactorsMicroreactorcontainscompleteamplificationmixC)AmplifyD)BreakMicroreactorsE)EnrichforDNApositiveBeforePCRAfterPCRGSFLX+ChemistryEmulsionPCRBasesareflowedsequentiallyandinaspecifiedcycleorder(TACG)AnucleotidecomplementarytothetemplatestrandgenerateslightThelightsignalisrecordedbytheCCDcameraduringeachflowSignalintensityisproportionaltothenumberofnucleotidesincorporatedMassivelyparallelsequencingofclonallyamplifiedbeadsinpicoliter-sizewells454SequencingSystemsKeyConceptwww.
454.
comSequencingBySynthesisAATCGGCATGCTAAAACTGARepeateddNTPflowsequence:AAGCTAAnnealPrimerProcesscontinuesuntiluser-definednumberofnucleotideflowcyclesarecompleted.
GSFLX+ChemistrySequencing-by-synthesisGCTATTTGCCGCGACGTTTTTADataProcessingGSRunProcessorTAGCTImageProcessingRegiondefinition&backgroundsubtractionWellidentificationonthePTPdeviceSignalProcessingNormalization&correctionstepsQualityfiltering&qualityreadtrimmingFlowgramgeneration&basecallingTACGFlowOrderKeysequenceTCAGorGACT(libraryreads)KeysequenceCATGorATGC(controlreads)UsedforwellidentificationandsignalcalibrationTTCTGCGAA454SequencingSystemsFlowgramGeneration&Basecallingwww.
454.
comDataAnalysisOverview(SFFTools)–SFFfilemanipulationtoolsGSdenovoAssembler–GenomeorTranscriptomedenovoassembly;Largegenomesuptohumansize;Hybridassemblies–Pairwisealignmentsofreadsresultinginconsensusandcorrespondingqualityscoresofcontigs,ACEfileofthealignment,metricsfilesandcontigscaffolds(whenusingPaired-enddata)GSReferenceMapper–TargetedmappingusingSequenceCapture;GenomeorTranscriptomereferencemapping;Variationdetectionincludinglargerearrangements–Individualreadsmappedtoareferencesequenceresultinginconsensusandcorrespondingqualityscoreofcontigs,ACEfileofthealignment,metricsfilesandlistofvariants(differencebetweenthesampleconsensusandreference)andstructuralrearrangements.
GSAmpliconVariantAnalyzer–Ultra-deepsequencingforvariantanalyses;Haplotyping–IdentificationandquantitationofsequencevariantsinanWhyReadLength,Speed&QualityMatterSequencingakillerE.
colibugwww.
454.
comDeadlyE.
colioutbreakinEuropebeganinMay2011ScientistsfromvariousinstitutesandcompaniesaroundtheglobequicklysequencedstrainisolatesThosewithfirstaccesstosampleshadtheleadwithspeed,butwhatofdataqualityandreadlengthWiththeGSJuniorassembly,theexpertswereabletoanswerbiologically-relevantquestionswhereotherassembliesfailed.
www.
454.
comE.
coliO104OutbreakSequencingtimelineMay24June1June2June6June10Germanauthoritiesreportthreedeathsand80casesofhaemolyticuraemicsyndromefromE.
coliinfectionEurope'stophealthofficialannounces"aseriouscrisis"UKHealthProtectionAgencyreleasesdenovoassemblyofH112180280isolateusingGSJuniorSystemUniversityHospitalMuenster&IonTorrentannounce"assembly"ofLB226692isolateBeijingGenomicsInstitutereleasesassemblyofTY2428isolate10PGMruns7PGM+1HiSeqruns3GSJuniorrunswww.
454.
comE.
coliO104OutbreakSequencingtimelineMay24June1June2June9June10"Moresequencedata,particularlypaired-endsequencingandlongerreadsthanwehaveinthecurrentassemblyarereallyneededtoresolvetheseprophageregions.
.
.
"NicoPetty,UniversityofQueenslandCommunityembracesHPAassemblyasreference–answersbiologicallymeaningfulquestionsBioinformaticsCommunityHitsaWallSource:http://bacpathgenomics.
wordpress.
com/2011/06/09/german-e-coli-phage-analysis-by-nico-petty/Highqualitydraftassembly(13scaffolds)Lotsofquestions……answers!
Globalbioinformaticscommunityanalyzingshort-readdatasetsLotsofdatabutlimitedcontextHighlyfragmentedassembliesrangingfrom>450to>3,000contigsRecognizedoutbreakstraingenomewasdifferentbutcouldn'tarrangethepieces"Afteronly5minutesoflookingovertheHPAassemblyweveryclearlysawwhatwehadexpectedtoseeallalongbutwasmissinginthepreviousassemblies–a62KbphageinsertioneventinthewrbAgene.
"DavidHirschberg,ColumbiaUniversityBGIReleasesIonTorrent–IlluminaAssemblyLotsofdatabutfewanswersBGIAnalysisWorkflowwww.
454.
comComplicatedpost-assemblyanalysis(evenfortheexperts)Complex,manualassemblyprocessQuestionsremainWhyisthisstrainsotoxicSomepiecesdon'tmatchtheexpectedbiology!
(Isthatreal)Source:http://bacpathgenomics.
wordpress.
com/2011/06/09/german-e-coli-phage-analysis-by-nico-petty/HPAReleasesGSJuniorAssemblyCommunitystartsansweringquestionswww.
454.
comGSJuniorSequenceDataGeneratedDeNovoAssembly(GSAssembler)Push-buttondenovoassemblyPerformedbyhealthcarescientistatHPA–notabioinformatician!
Nowthestorystartstoform…LargeinsertioneventsAcquiredmulti-drugresistanceAdhesin(knownvirulencefactor)Integratedintothechromosome!
Source:http://bacpathgenomics.
wordpress.
com/2011/06/11/tn21-resistance-transposon-in-the-chromosome454WorkflowThebiologybecomesclear~1minuteset-up~20mincomputationAnswersinhoursStx-phagescarriesgenesforShigatoxinPhage(62kblong)istypicallyintegratedintothewrbAgeneBothshortreadassembliesfailedtoaccuratelyreconstructthisportionofthegenomeRepresentationofStx2phageinsertiononHPAGSJuniorscaffold1WrbA5'endWrbA3'endStx2convertingphageTheBiology:WhyLongQualityReadsMatterBiologicallyaccurateinsertionreconstructionGSJuniorassemblycorrectlycontainstheinterruptedwrbAgenewithstx2phageinsertedonasinglescaffoldasisexpectedfromthebiologyBlogTitle:"Readlengthmatters:identifyingthephiStx2attsite""AmultisequencealignmentofthewrbAgeneclearlyrevealsthattheLife-TechnologiesAFOB00000000assemblingisnotreliableattheattachmentsite.
ItcontainsthreecontigsmatchingthewrbAgene,twoofthem(AFOB01000188.
1,AFOB01000143.
1)representingtheinterruptedgene,whilethethirdcontig(AFOB01000030.
1)showsanuninterruptedgene(thoughwithadistortedreadingframe).
"AssemblyComparisonOverviewFewerruns,morecompleteassembly,betterbiologyHPAAssemblyvs.
BGIAssembly3GSJuniorruns7IonTorrent+unspecifiedHiSeqrunsSingle-stepassemblyComplexmanualexpertassembly13scaffolds(35Xfewer)453scaffolds969KbN50scaffold(8.
5Xlonger)115KbN50scaffold~50%ofassembledbasesinscaffolds>1Mb0%Note:WithoutIlluminadata,IonTorrent&www.
454.
comCostComparisonOverviewFewerruns,easieranalysis,lowercostHPAGSJuniorassembly–3runs=~$3,000–1instrument=$100,000–Totalsequencingruntime=~1day–Assemblysoftware=FREE–Bioinformaticsexpertise=molecularbiologistfriendlyBGI(IonTorrent&Illumina)assembly–7IonTorrentruns=~$3,500–UnknownHiSeqpairedendruns=$20,000–1IonTorrentPGM&1IlluminaHiSeqinstrument=~$750,000–Totalsequencingruntime=1week+–AssemblySoftware/bioinformaticsexpertise=$$Expensive!
www.
454.
comGSJuniorAssemblyOverviewDeliveringwhatthecommunityneededUponrelease,communityimmediatelyembracedHPAassemblyasthereferenceGSJuniorassemblyprovidedclearanswerstokeybiologicalquestionsregardingthepathogenicityofthisuniquestrain–Correctlyidentifiedandlocateddrugresistanceandtoxingenesintegrationintochromosome–whileotherassembliescouldn't–Denovoassembledmajorityofplasmidcontent–Correctedmis-assembledphageregionsintheshort-readassembliesAfterworkingwithshortreadassembliesfordays,expertsturnedtotheGSJuniordataandwithin~5minutesfoundtheanswersPush-buttondenovoassemblyBiologicalanswers3GSJuniorRuns454SequencingDevelopmentsImprovingeaseofuse,sequencingperformance,andapplicationsUpfrontWorkflowAutomationSequencingReadLengthImprovementsApplications&AssaysEaseofuseimprovementstoGSJuniorworkflowStep-by-stepautomationsolutionsforlibraryprepandemPCR(breaking&enrichment)ExtendGSJuniorreadlengthbeyond400bp(shotgun&licon)OptimizeGSFLX+longreadsforampliconseqImproverobustnessandconsistencyofresultsLaunchdedicatedassaysforHLA,leukemiaandHIV/HBV/HCVPlannedCE-IVDapprovedkitsforselectassaysGSGTypeAssaysfor454SequencingSystemsAvailableNow!
GSGTypeLeukemiaPrimerSets(TET2/CBL/KRAS&RUNX1)GSGTypeHLAMR&HRPrimerSetsSequence-basedassaysforhigh-andmedium-resolutionHLAtypingForuseoneithertheGSJuniororGSFLXSystemSequence-basedassaysforleukemiaresearchForuseoneithertheGSJuniororGSFLXSystemExpanded454AssayMenuBeyond2012–focusonvirologyassaysVirology–GSVTectHIVResistance(PlannedCE-IVDinEurope)–HCVResistance/Genotyping(PlannedCE-IVDinEurope)–HBVResistance/Genotyping(PlannedCE-IVDinEurope)HLA–GSGTypeHLAPrimerSets(LSR)-AvailableNow!
–GSGTypeHLARegistry(PlannedCE-IVDinEurope)CollaborationwithDNAElectronicsISFETSequencingTechnologyCollaborationtodevelopsemiconductor-basedsequencingsystemLicenseDNAe'sISFET(ion-sensitive,fieldeffecttransistor)sequencingtechnologyEnablesdetectionofpHchangesfromnucleotideincorporationonsemiconductorchipBuildsoncurrent454SequencingportfoliobymovingfromopticaltoelectrochemicaldetectionTargetapplicationsinampliconassays,genepanels,exomesequencing,RNA-seq,andwholeDNAElectronicsSpunoutofImperialCollegeLondonDescriptionHigh-densityarrayofmicro-wellsonanintegratedcircuit.
EachwellholdsauniqueDNAtemplatebead.
Beneatheachwellisanion-sensitivetransistor.
Thechipissequentiallyfloodedwithonenucleotideafteranother.
IfthenucleotidethatfloodsthechipisnotamatchtotheDNAtemplate,nochangewillbedetectedandnobasewillbecalled.
IfanucleotideisincorporatedintotheCollaborationwithDNAe:TechnologyISFETsenablelabelfreepHSensingonaSemiconductorChipTechnologyConceptDescriptionMicrowellArrayPackagedIntegratedCircuitH+ionsdetectedCollaborationwithIBMNanoporeSequencingTechnologyIBMT.
J.
WatsonResearchCenterDescriptionCollaborationwithIBMResearchtodevelopnanopore-basedsequencingsystemTechnologybasedonIBMDNATransistorconceptLeverageIBMasaworldleaderinmicroelectronics,informationtechnologyandcomputationalbiologyAccesstoworldclassprototypefacilitiesforsemiconductormanufacturingandnanofabricationIBMDNAtransistorconceptcomprisesoflayersofinterdigitatedelectrodeswithdielectricmaterial,builtintoamembranethatcontainsthenanopores.
AnelectricfieldisusedtomoveDNAacrossananoporeonebaseatatime(electrophoreticratchet).
TheelectricfieldwithinthenanoporeinteractswithelectricchargesonDNAbackbonewhereindividualnucleotidesaretrappedatlocalfieldminima.
SwitchingthepolarityofthevoltageacrosstheelectrodespermitsthedriftoftheDNAmoleculeinonedirectionuntilthenucleotidesgettrappedatnewlocalfieldminimaAnucleotidesensingmethodologybasedonrecognitiontunnelingfromASUwillbecoupledtotheDNAtransistortodeterminetheCollaborationwithIBM:TechnologyConceptCombiningadielectricnanoporetransistorwithanucleotidesensingtechnologytoidentifyDNAbasesDNATransistorTechnologyConceptDescriptionNanoporeLocalElectricFieldMinimaDNAWatchthetechnologyvideoat:http://www.
youtube.
com/watchv=wvclP3GySUY454SequencingDevelopmentsSummaryOurobjectiveisto:improve–GSJunior&GSFLX+Systemsstrengthen-Assaysgrow–DNAecollaborationinnovate–IBMcollaborationwww.
454.
comForlifescienceresearchonly.
Notforuseindiagnosticprocedures.
454,454SEQUENCING,454LIFESCIENCES,EMPCR,GS20,GSFLX,GSJUNIOR,GSFLXTITANIUM,PTP,FASTSTARTandPICOTITERPLATEaretrademarksofRoche.
RocheTechnicalSupportus.
gssupport@roche.
com1.
800.
262.
49112-23-12Version1AnotherRecentExampleSequencingStreptococcusinToxicScarletFeverOutbreakJune15:UniversityofHongKongusedGSFLXSystemtosequenceStreptococcusstrainthatcausedscarletfeveroutbreakandonereporteddeathDraftgenomefinishedwithin3days–GSFLXrun(514,422reads)assembledusingGSDeNovoAssembler–47contigs–81KbN50contigsizeSource:http://www.
hku.
hk/press/news_detail_6505.
htmlResearchersidentifiedauniquemutation(48Kbinsertion)withingenomethatisabsentinotherpublishedgenomesofrelatedstrains
- coliwordpress安装相关文档
- wpwordpress安装
- NEYwordpress安装
- 英国wordpress安装
- 内容wordpress安装
- 开发wordpress安装
- 注册wordpress安装
RackNerd:便宜vps补货/1核/768M内存/12G SSD/2T流量/1G带宽,可选机房圣何塞/芝加哥/达拉斯/亚特拉大/荷兰/$9.49/年
RackNerd今天补货了3款便宜vps,最便宜的仅$9.49/年, 硬盘是SSD RAID-10 Storage,共享G口带宽,最低配给的流量也有2T,注意,这3款补货的便宜vps是intel平台。官方网站便宜VPS套餐机型均为KVM虚拟,SolusVM Control Panel ,硬盘是SSD RAID-10 Storage,共享G口带宽,大流量。CPU:1核心内存:768 MB硬盘:12 ...
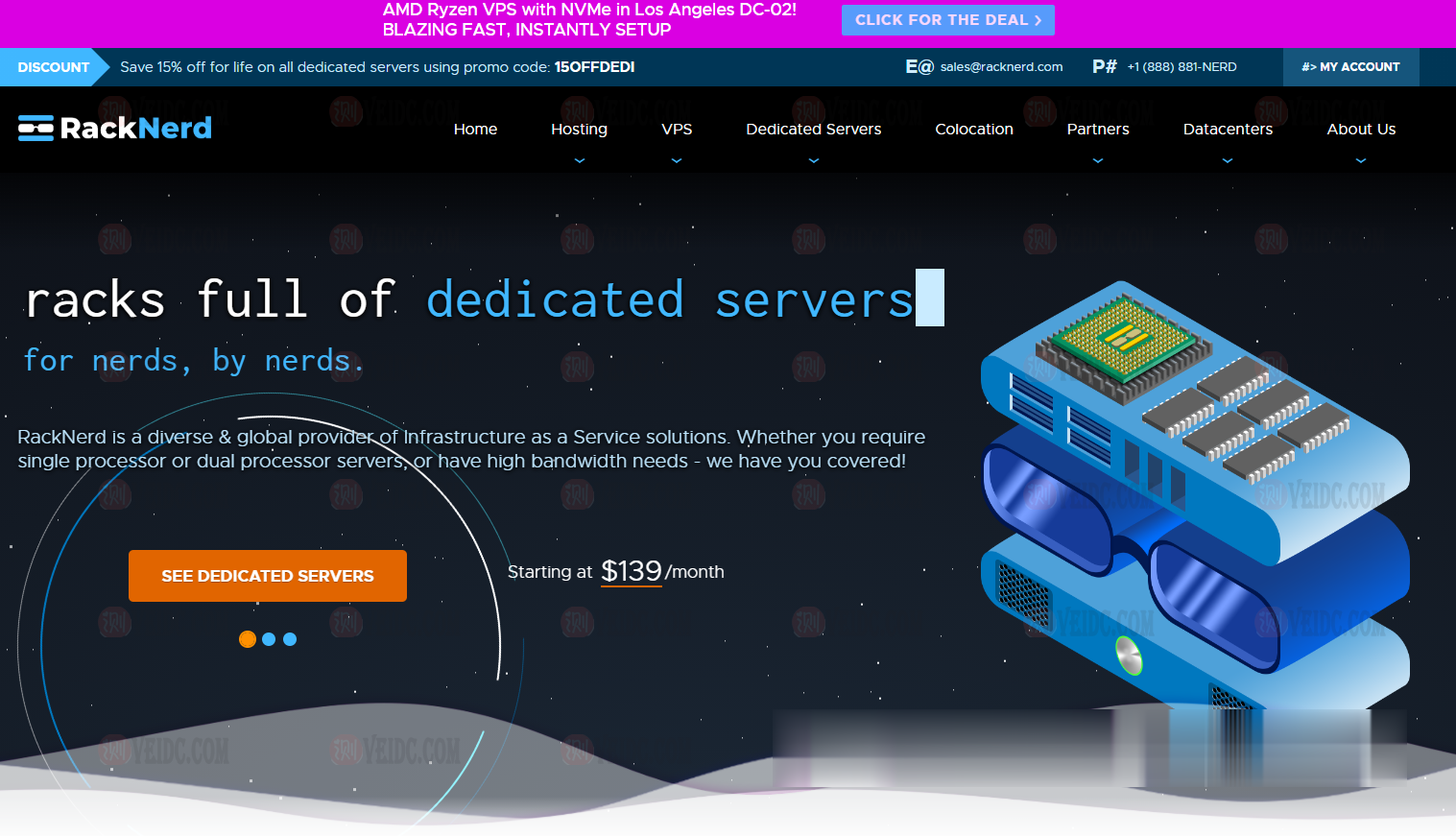
简单测评melbicom俄罗斯莫斯科数据中心的VPS,三网CN2回国,电信双程cn2
melbicom从2015年就开始运作了,在国内也是有一定的粉丝群,站长最早是从2017年开始介绍melbicom。上一次测评melbicom是在2018年,由于期间有不少人持续关注这个品牌,而且站长貌似也听说过路由什么的有变动的迹象。为此,今天重新对莫斯科数据中心的VPS进行一次简单测评,数据仅供参考。官方网站: https://melbicom.net比特币、信用卡、PayPal、支付宝、银联...
舍利云30元/月起;美国CERA云服务器,原生ip,低至28元/月起
目前舍利云服务器的主要特色是适合seo和建站,性价比方面非常不错,舍利云的产品以BGP线路速度优质稳定而著称,对于产品的线路和带宽有着极其严格的讲究,这主要表现在其对母鸡的超售有严格的管控,与此同时舍利云也尽心尽力为用户提供完美服务。目前,香港cn2云服务器,5M/10M带宽,价格低至30元/月,可试用1天;;美国cera云服务器,原生ip,低至28元/月起。一、香港CN2云服务器香港CN2精品线...
wordpress安装为你推荐
-
国际域名国际域名和国内域名有什么不同,什么叫顶级域名?英文域名英文域名与中文域名有啥区别免费国内空间网站免费空间(国内的)那里有?域名购买域名注册和购买是一个意思吗?美国vps租用如何租用到最快的美国服务器手机网站空间手机网页空间需要多大?香港虚拟主机想买一个香港虚拟主机,大家推荐一下吧虚拟主机评测网哪里有可靠的免费虚拟主机虚拟主机系统什么是虚拟主机?论坛虚拟主机论坛虚拟主机的IP地址在后台的那个地方呀